Distinction of solid liquid atoms and clustering¶
In this example, we will take one configuration from a molecular dynamics simulation which has a solid cluster in liquid. The task is to identify solid atoms and cluster them. More details about the method can be found here.
The first step is, of course, importing all the necessary module. For visualisation, we will use Ovito.

The above image shows a visualization of the system using Ovito. Importing modules,
import pyscal.core as pc
Now we will set up a System with this input file, and calculate
neighbors. Here we will use a cutoff method to find neighbors. More
details about finding neighbors can be found
here.
sys = pc.System()
sys.read_inputfile('cluster.dump')
sys.find_neighbors(method='cutoff', cutoff=3.63)
Once we compute the neighbors, the next step is to find solid atoms.
This can be done using find_solids() method.
sys.find_solids(bonds=6, threshold=0.5, avgthreshold=0.6, cluster=False)
The above statement found all the solid atoms. Solid atoms can be
identified by the value of the solid attribute. For that we first
get the atom objects and select those with solid value as True.
atoms = sys.atoms
solids = [atom for atom in atoms if atom.solid]
len(solids)
202
There are 202 solid atoms in the system. In order to visualise in Ovito,
we need to first write it out to a trajectory file. This can be done
with the help of to_file() method of System. This method can help to
save any attribute of the atom or any Steinhardt parameter value.
sys.to_file('sys.solid.dat', custom = ['solid'])
We can now visualize this file in Ovito. After opening the file in
Ovito, the modifier compute
property
can be selected. The Output property should be selection and in
the expression field, solid==0 can be selected to select all the non
solid atoms. Applying a modifier delete selected
particles
can be applied to delete all the non solid particles. The system after
removing all the liquid atoms is shown below.
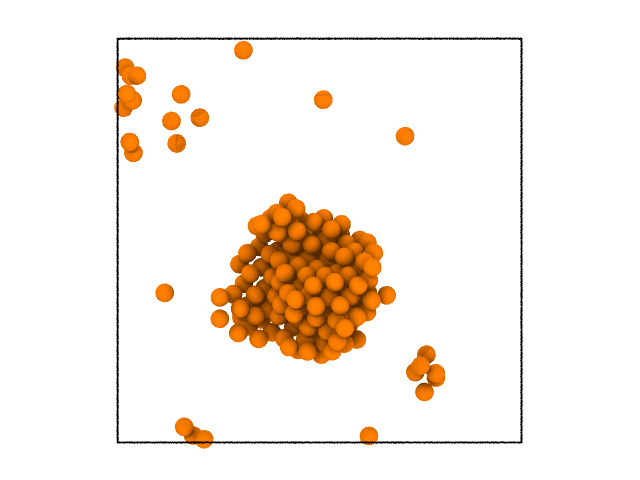
Clustering algorithm¶
You can see that there is a cluster of atom. The clustering functions
that pyscal offers helps to select atoms that belong to this cluster. If you used find_clusters()
with cluster=True, the clustering is carried out. Since we did used
cluster=False above, we will rerun the function
sys.find_solids(bonds=6, threshold=0.5, avgthreshold=0.6, cluster=True)
176
You can see that the above function call returned the number of atoms
belonging to the largest cluster as an output. In order to extract atoms
that belong to the largest cluster, we can use the largest_cluster
attribute of the atom.
atoms = sys.atoms
largest_cluster = [atom for atom in atoms if atom.largest_cluster]
len(largest_cluster)
176
The value matches that given by the function. Once again we will save this information to a file and visualize it in Ovito.
sys.to_file('sys.cluster.dat', custom = ['solid', 'largest_cluster'])
The system visualized in Ovito following similar steps as above is shown below.
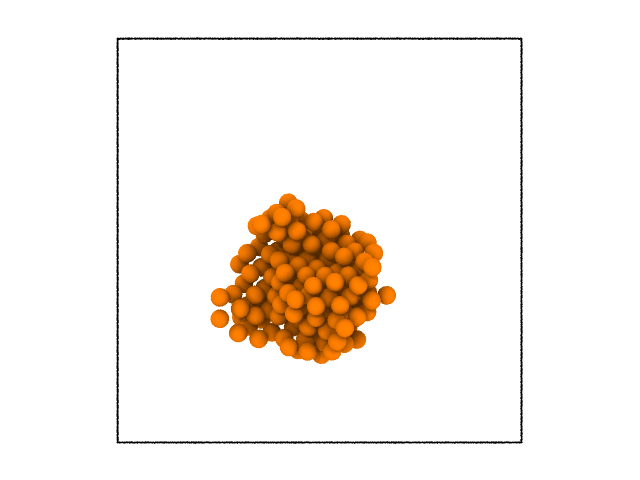

It is clear from the image that the largest cluster of solid atoms was successfully identified. Clustering can be done over any property. The following example with the same system will illustrate this.
Clustering based on a custom property¶
In pyscal, clustering can be done based on any property. The following
example illustrates this. To find the clusters based on a custom
property, the cluster_atoms() method has to be used. The
simulation box shown above has the centre roughly at (25, 25, 25). For
the custom clustering, we will cluster all atoms within a distance of 10
from the the rough centre of the box at (25, 25, 25). Let us define a
function that checks the above condition.
def check_distance(atom):
#get position of atom
pos = atom.pos
#calculate distance from (25, 25, 25)
dist = ((pos[0]-25)**2 + (pos[1]-25)**2 + (pos[2]-25)**2)**0.5
#check if dist < 10
return (dist <= 10)
The above function would return True or False depending on a condition and takes the Atom as an argument. These are the two important conditions to be satisfied. Now we can pass this function to cluster. First, set up the system and find the neighbors.
sys = pc.System()
sys.read_inputfile('cluster.dump')
sys.find_neighbors(method='cutoff', cutoff=3.63)
Now cluster
sys.cluster_atoms(check_distance)
242
There are 242 atoms in the cluster! Once again we can check this, save to a file and visualize in ovito.
atoms = sys.atoms
largest_cluster = [atom for atom in atoms if atom.largest_cluster]
len(largest_cluster)
242
sys.to_file('sys.dist.dat', custom = ['solid', 'largest_cluster'])
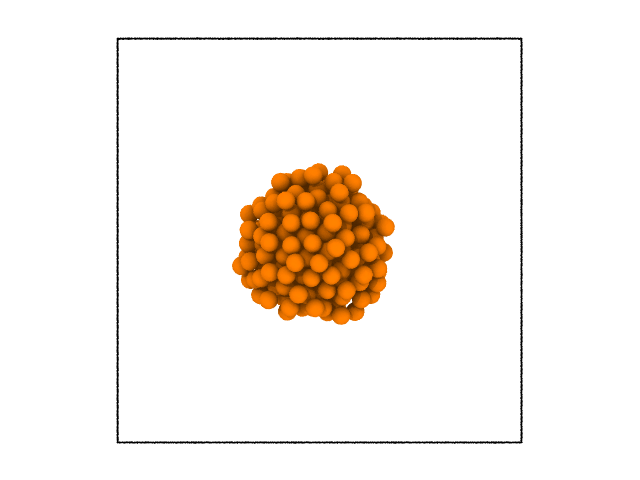

This example illustrates that any property can be used to cluster the atoms!